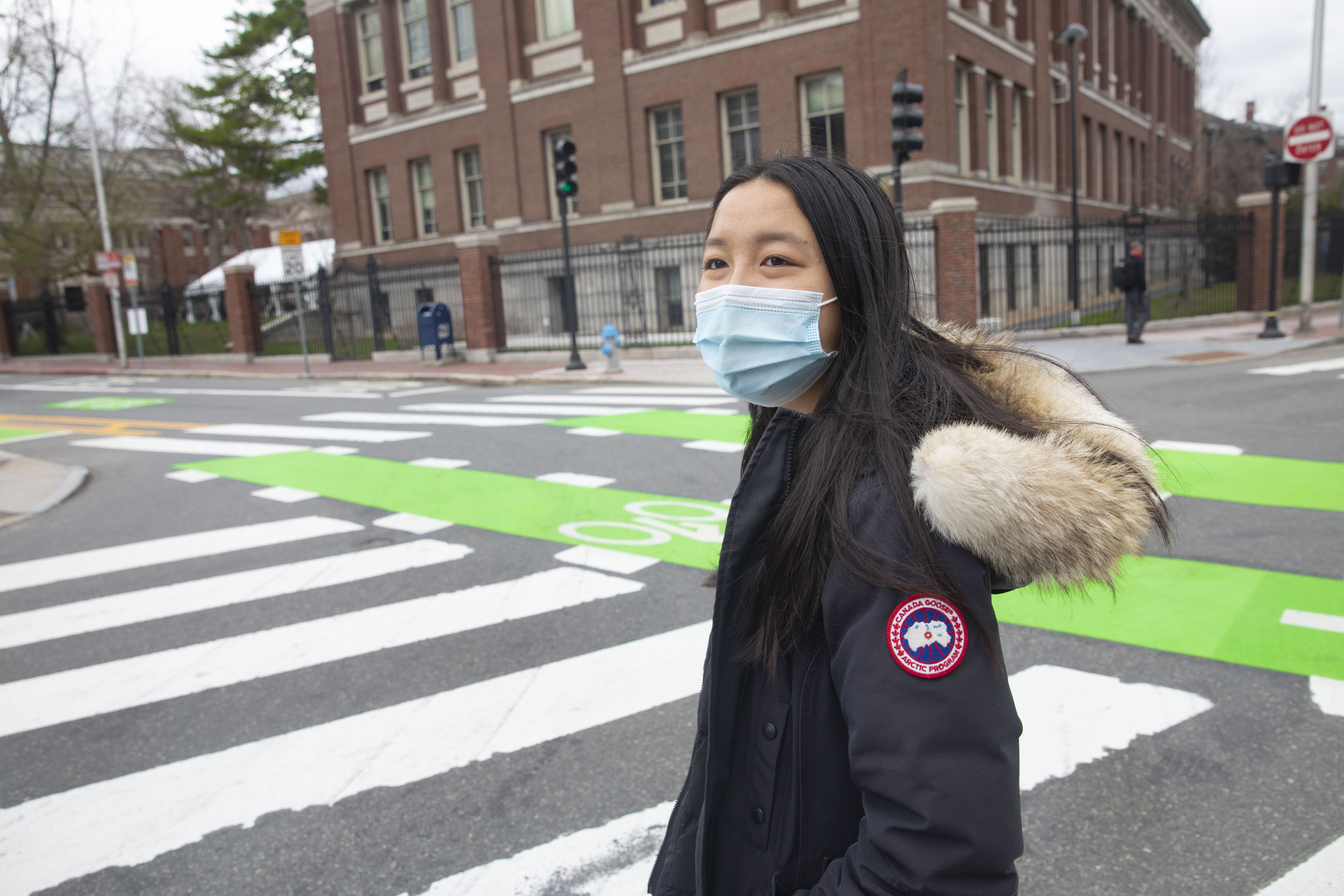
Tina Lu ’21 examined symptom internet searches early in the pandemic and scored how well they later matched clinical symptoms of COVID-19.
Kris Snibbe/Harvard Staff Photographer
Tracking progression of disease through internet searches for symptoms
Student’s project found queries mirrored course of illness, foretold rise in cases
You’re not feeling well so you open a search engine and type: fever, dry cough, hoping to find hints of what you may have. A handful of days later, you’re feeling worse, and you type in: trouble breathing. It turns out you’re not the only one who’s doing this, and a Harvard senior’s research project suggests that tracking the results of all those searches can tell us something about the progression of a new disease in individuals and through a population.
Tina Lu, a Leverett House computer science concentrator, analyzed search engine data from Google Trends going back to the beginning of the coronavirus pandemic to see how well symptom searches in 32 countries on six continents matched the clinical symptoms of COVID-19 and whether the number of searches served as a harbinger of rising incidence of cases.
“We hope that our findings that this is true for COVID will be helpful for future pandemics because it might take a while for research to be done on the course of illness for a new disease, but Google Trends can be analyzed from the very beginning,” Lu said.
Published in Nature Digital Medicine earlier this year, the work found that from Jan. 1, 2020, to April 20, 2020, increases in symptom-related searches for COVID-19 preceded increases in reported cases and deaths by an average of 18.53 days. Further, the work showed that there was a clear pattern of disease progression, with early symptoms of fever, dry cough, sore throat, and chills being followed by more severe shortness of breath by 5.22 days, matching the clinical course of disease reported in medical studies.
“This is not just a one-time contribution. It’s a new tool for the public health community,” said Ben Reis, assistant professor of pediatrics at Harvard Medical School, director of the Predictive Medicine Group at the Boston Children’s Hospital Computational Health Informatics Program, and senior author on the paper. “When you’re faced with a novel pathogen, the most valuable resource is information, especially in the early stages.”
One way that modern medicine comes to understand new ailments is through case reports published in medical journals. That was the case with COVID-19, Reis said, and those observations by trained physicians will remain a valuable source of information. But journal publication can be slow, Reis said, while search engine data like that available through Google Trends can be gathered the following day or even the same day, offering a fast snapshot of what patients are experiencing and how their disease is progressing. In the case of a rapidly-moving pathogen like SARS-CoV-2, the digital data has the potential to provide invaluable early insights, Reis said.
“When a novel pandemic strikes, in addition to the existing established approaches, we propose that this could be a complementary approach to start looking at the data for different symptoms and symptom groups in any area that’s affected,” Reis said. “We’re offering an additional, complementary data source to existing sources that has the benefit of very quick availability. Google Trends or any other search engine data is available in near real time, you can get data about yesterday and in some cases, even about today.”
“We hope that our findings … will be helpful for future pandemics because it might take a while for research to be done on the course of illness for a new disease, but Google Trends can be analyzed from the very beginning.”
Tina Lu
The project got underway in the months before the pandemic when Lu, then a junior, reached out to Reis at his lab at Boston Children’s Hospital. Lu was interested in applying her computer science skills to medical research and knew Reis’ lab had used Google Trends data for public health studies in the past, tracking data related to polio vaccination, for example.
The project, much of which Lu conducted from her home in New Jersey, was initially aimed at seeing whether disease peaks in different countries could be detected, but the work was confounded by different precautionary steps taken in different nations. Then she began looking at symptoms and noticed different symptoms peaked at different times in different countries. Lu looked closer and saw a clear pattern that matched what is now known as the clinical course of COVID-19.
“It was very exciting and also a little hard to completely believe at first, because I was investigating this for a really long time,” Lu said.
Lu, who was working out of her family’s basement, spent several months gathering and examining the data during the pandemic’s initial lockdown, while also juggling classwork from courses that had moved abruptly online in March 2020.
More like this
“I was spending hours each day for many days,” Lu said. “At first, it felt like I was not seeing any consistent patterns between different countries. I would have daily calls with Ben, and we would go over different approaches or different things to look into. When we found this pattern with the course of illness, it was just very exciting because finally we found something that I guess no one else had found before and that was pretty consistent across all of these different countries we were looking at.”
Lu, who graduates this spring, plans to head to Chicago after Commencement for a job as a software engineer with an investment firm headquartered there. Though the project is over, Reis said he expects it to have a real-world impact — potentially even on variants that emerge in this pandemic. The publication outlines the methodology so others can emulate it, and he’s already fielded several inquiries from researchers.
“We’re happy to work with others. The paper was written in a way so that anyone who’s working in this space could recreate it and incorporate these approaches,” Reis said. “Our hope is that these emerging systems that have been built over the last decade to incorporate digital information into the epidemiological information cycle will incorporate these methods as well.”