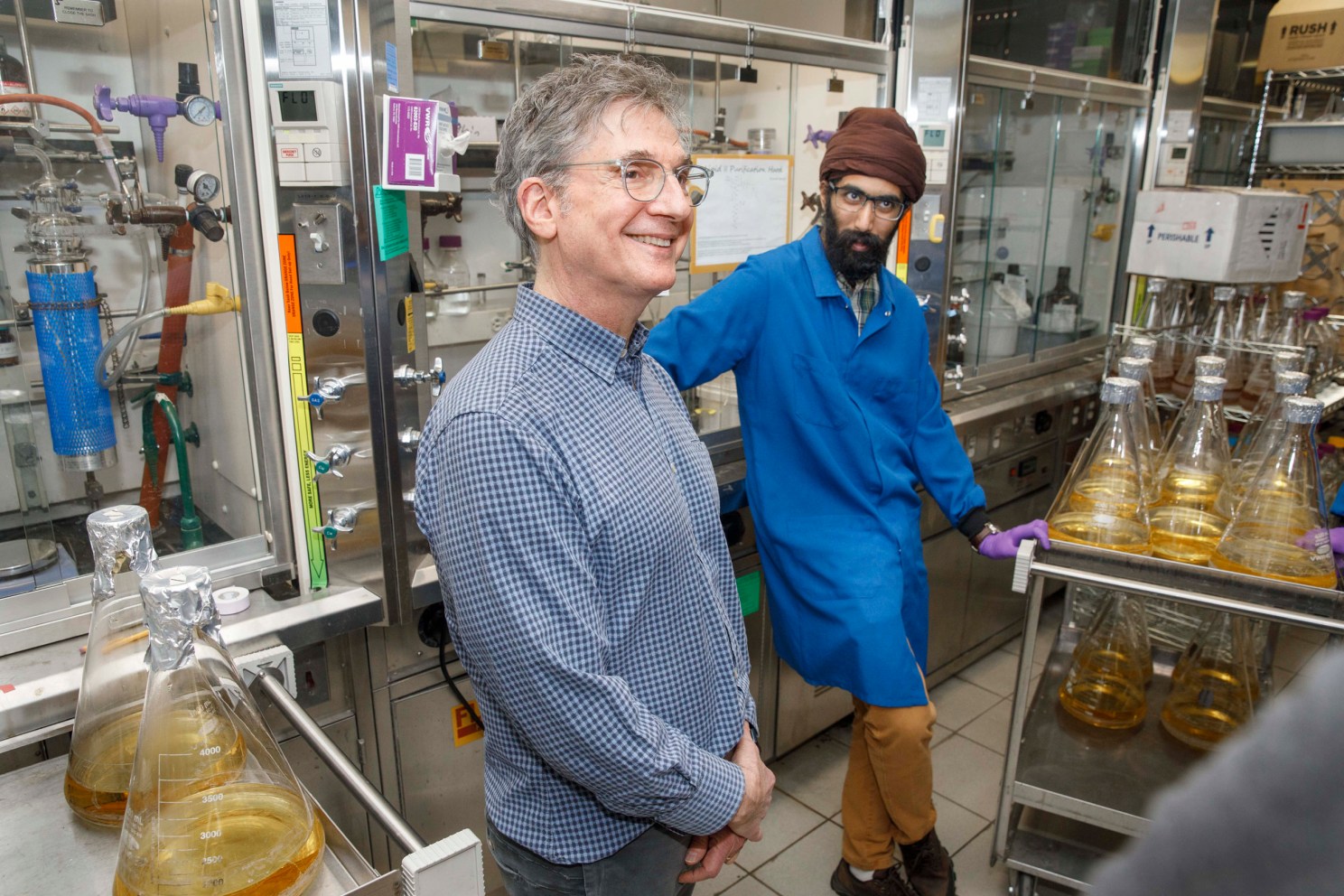
Co-authors Daniel Kahne and Karanbir Pahil in the lab.
Photos by Kris Snibbe/Harvard Staff Photographer
Finding flaws in superbugs’ armor
Harvard biochemists’ research paves the way for development of new antibiotics
Recent years have seen the rise of bacterial pathogens that have developed resistance to antibiotics. One such superbug, carbapenem-resistant Acinetobacter baumannii (CRAB), kills hundreds of critically ill patients in the U.S. each year, usually in hospital settings, by causing blood, lung, or urinary tract infections that don’t respond to treatments.
Daniel Kahne, Higgins Professor of Chemistry and Chemical Biology, has spent much of his career studying bacterial physiology, uncovering fundamental mechanisms by which these species thrive and evade attack.
The compound, called Zosurabalpin, is now in Phase I clinical trials. If approved, it would be the first new treatment for A. baumannii infections in 50 years.
His lab has a special interest in Gram-negative bacteria, which include A. baumannii and are characterized by an impermeable outer membrane that many drugs cannot cross. His team’s work is helping usher in new medications to combat this superbug, and perhaps others.
“For the last 25 years, I’ve been interested in how this outer membrane is constructed,” Kahne said. “There are a set of protein machines that are conserved in all Gram-negative bacteria that make this membrane, and so we study each of these machines.”
Kahne’s research has been key to the efforts of Roche, an international biotechnology company that recently announced a promising clinical candidate effective against the CRAB superbug. The compound, called Zosurabalpin, is now in Phase I clinical trials. If approved, it would be the first new treatment for A. baumannii infections in 50 years, and could offer a new weapon in the fight against antibiotic resistance.
The Gram-negative membranes Kahne studies are dotted with large, complex glycolipids called lipopolysaccharides (LPS), which act as protective armor. Bacteria assemble LPS molecules inside their cytoplasm before moving them to their outer membranes.
How bacteria orchestrate the transport of LPS from factory to final destination is what captivated Kahne. Scientists had long suspected that this multistep process could provide new targets for future antibiotic drugs.
Supported by the National Institutes of Health, in 2010 Kahne’s team was the first to propose a mechanism for how Gram-negative bacteria manage the transport. Describing a “trans-envelope bridge” made of seven different proteins, the researchers showed that LPS molecules use this bridge to travel from the cytoplasm, across an inner membrane, through an aqueous space called the periplasmic compartment, and across another membrane. Then, like a PEZ dispenser, the cell pushes LPS molecules out one at a time, where they line up on the cell’s surface.
The team published several more studies supporting their bridge model, showing in 2018 that they could reconstitute the seven-protein bridge from pure proteins in vitro. More recently, they reported using single-molecule tracking to show the bridges forming and transporting LPS in live cells.
Meanwhile, Roche had been working to develop a new type of antibiotic effective against CRAB infections. They had genetic evidence to believe their drug kills A. baumanni by disrupting the LPS transport machinery that Kahne’s lab has extensively studied, and which no other drug on the market currently has as its “lethal target.” In late 2020, Roche approached the Harvard researchers to help them make sure.
“Dan is a well-recognized world leader in this field, and we saw the potential for synergy between the two teams in order to validate the target and mechanism of action of Zosurabalpin as well as gain further insight into the fundamental biological process of LPS transport,” said Kenneth Bradley, head of infectious diseases discovery at Roche Pharma Research and Early Development.
With Roche scientists and collaborator Andrew Kruse, professor in biological chemistry and molecular pharmacology at Harvard Medical School, Kahne and team recently published a pair of Nature studies reporting on Roche’s promising new class of antibiotics in clinical development. One of the papers provided details of the drug’s LPS transport-based molecular mechanism.
“It was very exciting to be able to use all of the tools that the students in my group have developed over the last two decades to quickly figure out the mechanism of action of Roche’s compound,” Kahne said.
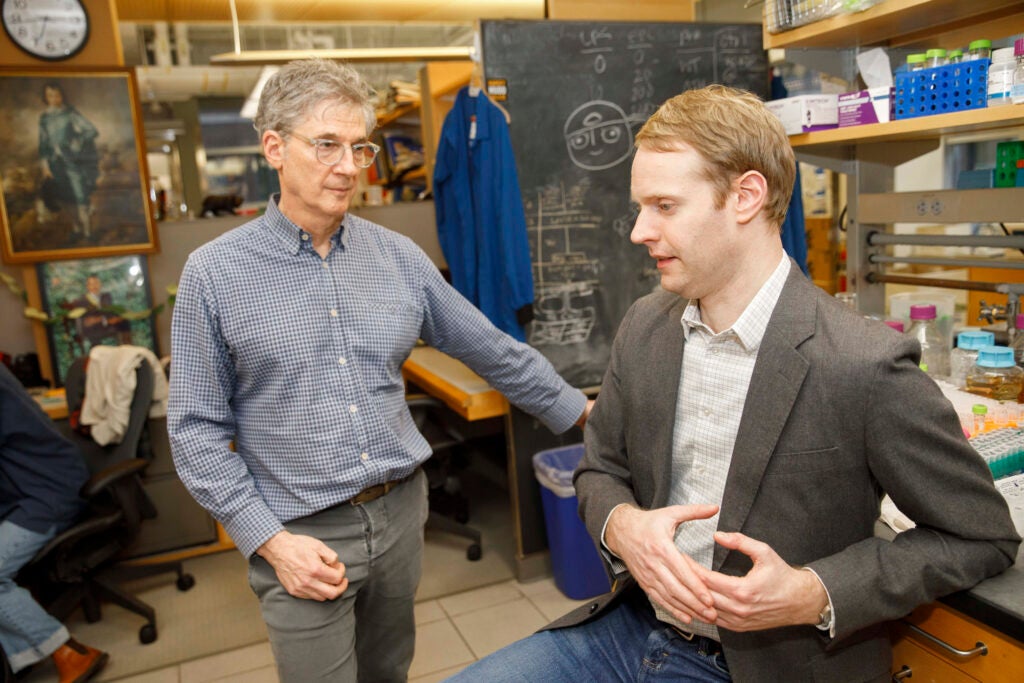

The tools included structural, biochemical, and genetic approaches to solve confounding protein structures. Karanbir Pahil, Ph.D. ’22, developed a lab procedure for producing large amounts of pure Acinetobacter transport proteins, whose complexity makes them difficult to work with. The research uncovered the first direct evidence that the Roche drug interferes with these complexes, killing the bacteria.
To capture the molecular details, they used cryo-electron microscopy, in which Kruse’s group has expertise. The technique is especially useful for studying intrinsically dynamic proteins with multiple conformations. “This very complicated transporter clearly falls into that category,” Kruse said.
Using Pahil’s lab-purified LPS transporter proteins, Morgan Gilman, a postdoctoral fellow in Kruse’s lab, taught Pahil how to generate and process cryo-electron microscopy data with the goal of determining the transporter proteins’ structure as it binds to Zosurabalpin.
Combining efforts, the Kahne and Kruse labs showed that the Roche drug candidate disrupts the transport of LPS to the cell surface by binding to a pocket that encompasses part of the protein bridge as well as the LPS molecule itself. The drug causes LPS to accumulate in the wrong part of the cell, “jamming” the LPS inside of its transporter, weakening the cell’s membrane, and eventually destroying the cell.
To bolster the evidence, Pahil, Gilman, and team solved a total of seven protein structures. “It required extensive optimization of expression and purification conditions to get well-behaved proteins,” Pahil said. Drugs that bind to a composite surface created by a protein machine and the cellular molecule it acts on are unusual, the researchers noted.
Zosurabalpin, if approved by the U.S. Food and Drug Administration, would be the first antibiotic to target the LPS transport mechanism. By contrast, most widely used antibiotics, including penicillin and carbapenem, are called beta-lactam antibiotics and work by inhibiting synthesis of the bacterial cell wall. Other types disrupt ribosomes and protein synthesis.
The pocket the Roche candidate drug binds to is specific to A. baumannii. But according to the researchers, a similar pocket exists in other Gram-negative bacteria, including E. coli, so they are excited by the possibility that the drug could be modified to treat other pathogens.
“We are starting to have a fairly deep understanding of how many of these molecular machines work,” Kruse said. “I think that’s going to be incredibly helpful in designing next-generation antibiotics.”
The Harvard Office of Technology Development spearheaded the three-year sponsored research agreement between Roche and Kahne’s lab that resulted in the pair of Nature papers.