A new approach to kill superbugs
An illustration of the Gram-negative bacteria’s red tint following a Gram-stain test.
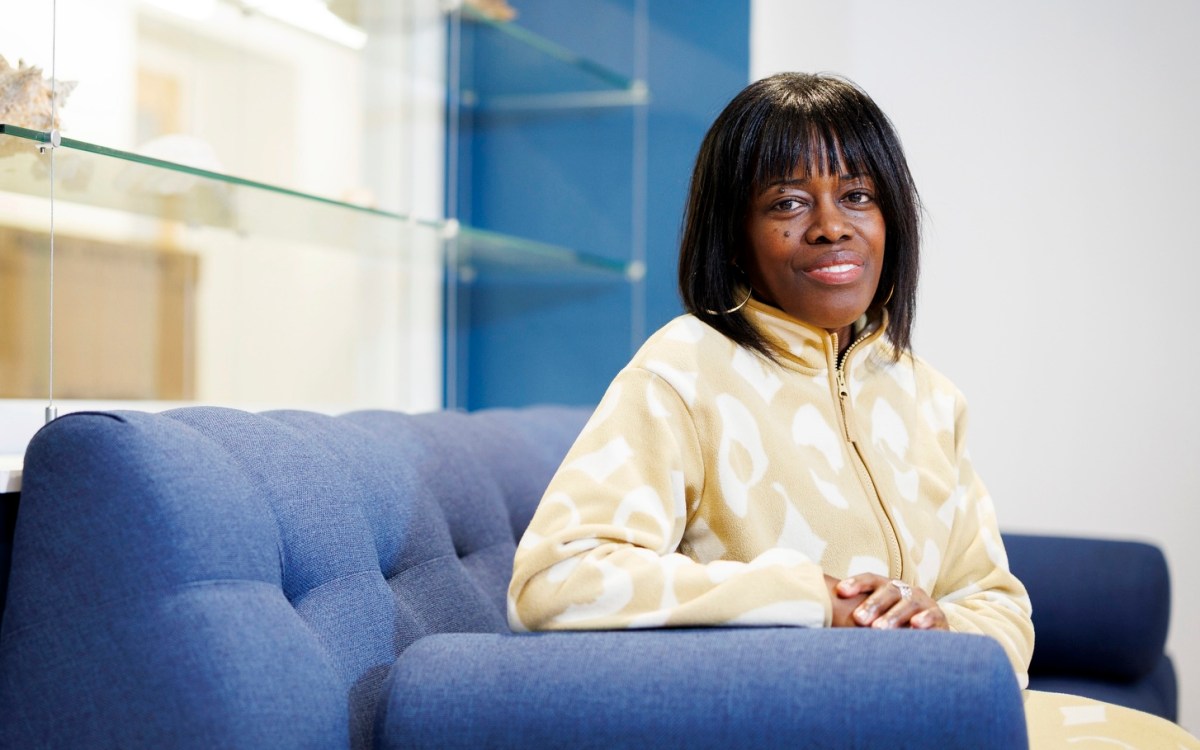
Daniel Kahne discovers how Gram-negative bacteria build resistance to antibiotics
In the late 1800s, bacteriologist Hans Christian Gram discovered a way to classify bacteria, most of which fall into one of two groups: Gram-positive or Gram-negative. He developed a stain, still in use today, which bacteria either absorb or repel. Gram-positive, with their adsorbent cell wall, glow violet, flush with dye. But Gram-negative bacteria boast an extra outer membrane that resists the stain. They remain a pointed pink or red.
This simple stain test reveals a complicated, pressing problem. Gram-negative bacteria, which cause a variety of life-threatening infections, develop greater antibiotic resistance every year. According to the National Institutes of Health, there is “increasing concern about drug resistance,” with some so-called superbugs “now resistant to many, most, or all available treatments.”
[gz_photo_layout_article_width image=”238869″ caption=”The%20many%20layers%20of%20a%20Gram-negative%20bacterium.” credit=”Credit%3A%20Kahne%20Lab” /]
For example, E. coli earns frequent headlines for food-borne outbreaks, other strains lead to pneumonia or bloodstream infections, and the infamous plague resulted from Yersinia pestis, yet another type of Gram-negative bacteria. In the last fifty years, no new drugs have been developed to combat this bacterial threat.
Now, a Harvard University team has discovered a new approach to address what Daniel Kahne calls a “major unmet clinical need.”
The team, which includes Kahne, the Higgins Professor of Chemistry and Chemical Biology, postdoctoral fellow Ran Xie, and graduate student Rebecca J. Taylor, authored a paper published in today’s Science Magazine. In it, Kahne and his team demystify a millennia-old process: how Gram-negative bacteria build their intractable outer membrane.
The membrane is fortified with the large glycolipid lipopolysaccharide (LPS). But, the mechanism by which the bacteria transport such a cumbersome lipid across multiple solid membranes remained a mystery.
“There have been two competing models in the field,” explained Rebecca. “This paper includes much better evidence for the second model.” The first (chaperone model) requires multiple molecular escorts to heave LPS from the energy-rich cell interior, or cytoplasm, across the energy-vacant membranes.
In the second model, weighty LPS molecules “are pushed one after the other in a PEZ dispenser-like manner across a protein bridge that connects the inner and outer membranes.” LPS molecules are loaded into the machine in the cytoplasm and “each new LPS molecule then pushes molecules already in the bridge toward the outer membrane.”
[gz_pull_quote attribution=”— Daniel Kahne, Higgins Professor of Chemistry and Chemical Biology”]“We answered a longstanding question in prokaryotic biology about how the outer membrane of Gram-negative bacteria is constructed. Hopefully these studies will help us identify new targets to treat resistant Gram-negative infections.”[/gz_pull_quote]
Already, Kahne’s team has demonstrated that interfering with membrane assembly makes Gram-negative bacteria “leaky” and vulnerable. Prior to this work, researchers had “no tools to study this process,” according to Kahne.
Understanding these molecular PEZ dispensers could help reverse this alarming healthcare crisis and, for the first time in fifty years, lead to new treatments for Gram-negative bacterial infections.




