The microbiomes in our gastrointestinal tracts regulate wide-ranging physiological, metabolic, immunologic, cognitive, behavioral, and psychiatric traits. Understanding and manipulating human microbiomes could be key to managing physical and mental health.
Credit: Wyss Institute at Harvard University
Microbiomes could hold keys to improving life
Newly formed consortium wants to accelerate microbiome research
Forty-eight scientists from 50 institutions in the U.S. have formed the Unified Microbiome Initiative Consortium (UMIC). The scientists envision a coordinated effort spanning national cross-institutional and cross-governmental agency support with the goal of driving forward cutting-edge microbiome research, enabling breakthrough advances in medicine, ecosystem management, sustainable energy, and production of commodities. Their proposal was published online in the journal Science on Oct. 28.
Microbial life forms — including viruses, bacteria, and fungi — are the most diverse and abundant organisms on earth. They have shaped our evolutionary origins for billions of years and continue to have widespread impact. The UMIC foresees the microbiomes being leveraged through genetic engineering for applications within 10 years.
“Microbes are everywhere. Therefore understanding microbiomes, whether they be the ones that live in and on our bodies or the ones in the environment, is essential to understanding life,” said Pamela Silver, a core faculty member at the Wyss Institute for Biologically Inspired Engineering at Harvard University.
Silver is also one of the faculty leaders on the Wyss Institute’s Synthetic Biology platform, the Elliot T. and Onie H. Adams Professor of Biochemistry and Systems Biology at Harvard Medical School (HMS), and a founding member of the Department of Systems Biology at HMS.
“Understanding how [microbiomes] work might hold the key to advances as diverse as fighting antibiotic resistance and autoimmune diseases, reclaiming ravaged farmland, reducing fertilizer and pesticide use, and converting sunlight into useful chemicals,” said Jeff F. Miller, director of the California NanoSystems Institute and corresponding author of the Science paper.
By metabolic processes, microbes synthesize countless different molecules, which through genetic engineering could lead to colonies of microbial “workers” being used for the sustainable synthesis of pharmaceuticals, materials, and chemical commodities. Genetically engineered microbes could also produce biofuels through metabolic processes and be used in the conversion of solar energy into liquid fuel, according to work already underway by Silver at the Wyss Institute and HMS.
Microbes also play a vital role in balancing biogeochemical processes, such as removing carbon dioxide from the atmosphere. The interactions between soil, plant roots, and microbes play an important role in plant health and crop yield.
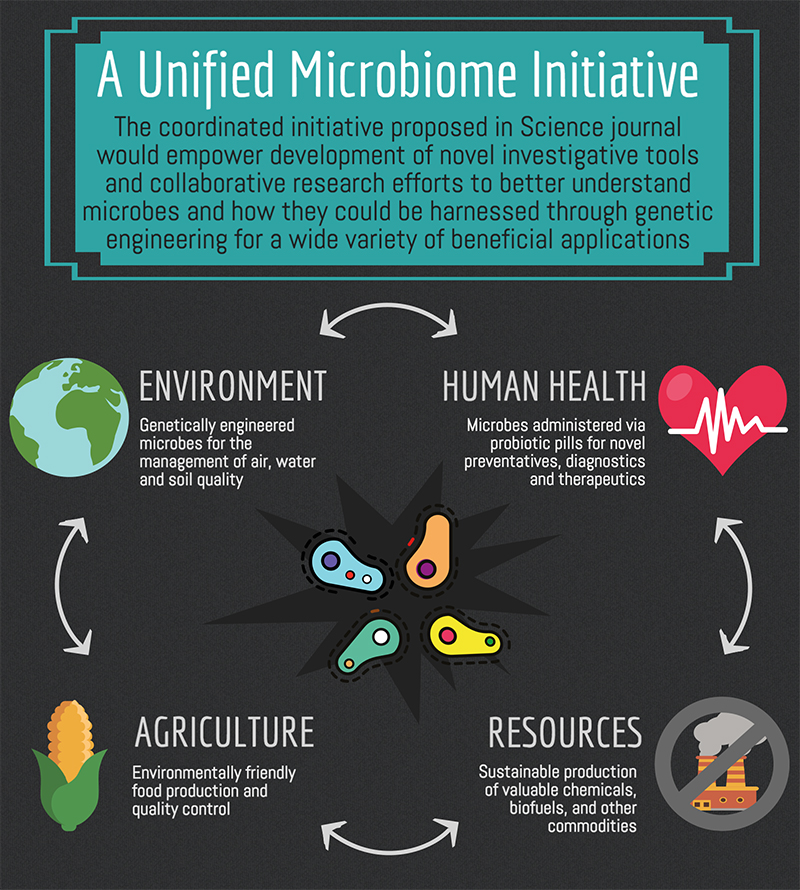

Furthermore, the microbiomes in our gastrointestinal tracts regulate wide-ranging physiological, metabolic, immunologic, cognitive, behavioral, and psychiatric traits. Understanding and manipulating human microbiomes could be key to managing physical and mental health. Silver and her team have already begun developing several potential avenues for leveraging gut microbes to improve health. In collaboration with Wyss core faculty member James Collins, Silver has engineered genetically programmed bacterial “reporters” that can detect and record conditions in the gastrointestinal tract. And, working with Donald Ingber, the Wyss Institute’s founding director, and its senior staff scientist Jeffrey Way, Silver is developing a consortia of synthetic microbes that could be used to treat gastrointestinal illness.
Her expertise and experience in these emerging areas of synthetic biology have enabled Silver to contribute her thought leadership as a member of the UMIC to how the proposed Unified Microbiome Initiative could integrate focus areas to accelerate microbiome research.
“I’m interested in engineering microbes as a way to interrogate their behavior,” said Silver. “The purpose of this unified initiative is to determine what are the big questions we have about the microbiome and what are the specific technologies we need in order to investigate those questions.”
Some of the big questions the group hopes to address through the organized coalition include understanding how microbes assemble into communities and what makes them resilient or resistant to perturbation, how genes in the microbiome interact with one another, which genes in the microbiome are associated with which organisms, as well as how we can beneficially harness the microbiomes of humans, animals, plants, and environments.
To find the answers to these questions, scientists must first be supported in the development of breakthrough technologies for investigating microbiomes. Specifically, the group recommends development of improved computational methods for analyzing and predicting the vast number of unknown genes and their functions comprising microbiomes; a transition from gene-specific to whole-genome-based analysis through improved genome reference libraries and sequencing methods; development of high-powered imaging methods for visually interrogating communities of microbes down to the individual level; new adaptive modeling systems and data reporting tools; improved genetic engineering techniques for perturbing microbial communities; and novel methods to mimic natural environments for supporting microbiome growth in the laboratory, among others.
“Discovery of the existence and importance of the microbiome has provided a new frame of reference for our understanding of health and our environment,” said Ingber, the Judah Folkman Professor of Vascular Biology at HMS and Boston Children’s Hospital. “A nationwide coordinated effort to invest in understanding and leveraging microbiomes could open entirely new frontiers in biotechnology and medicine, and lead to solutions that would not be possible in any other way.”
The paper calling for a Unified Microbiome Initiative can be found in the Oct. 30 issue of Science or through an online portal made available by the American Society for Microbiology.