Tracking entwined histories of malaria, humans
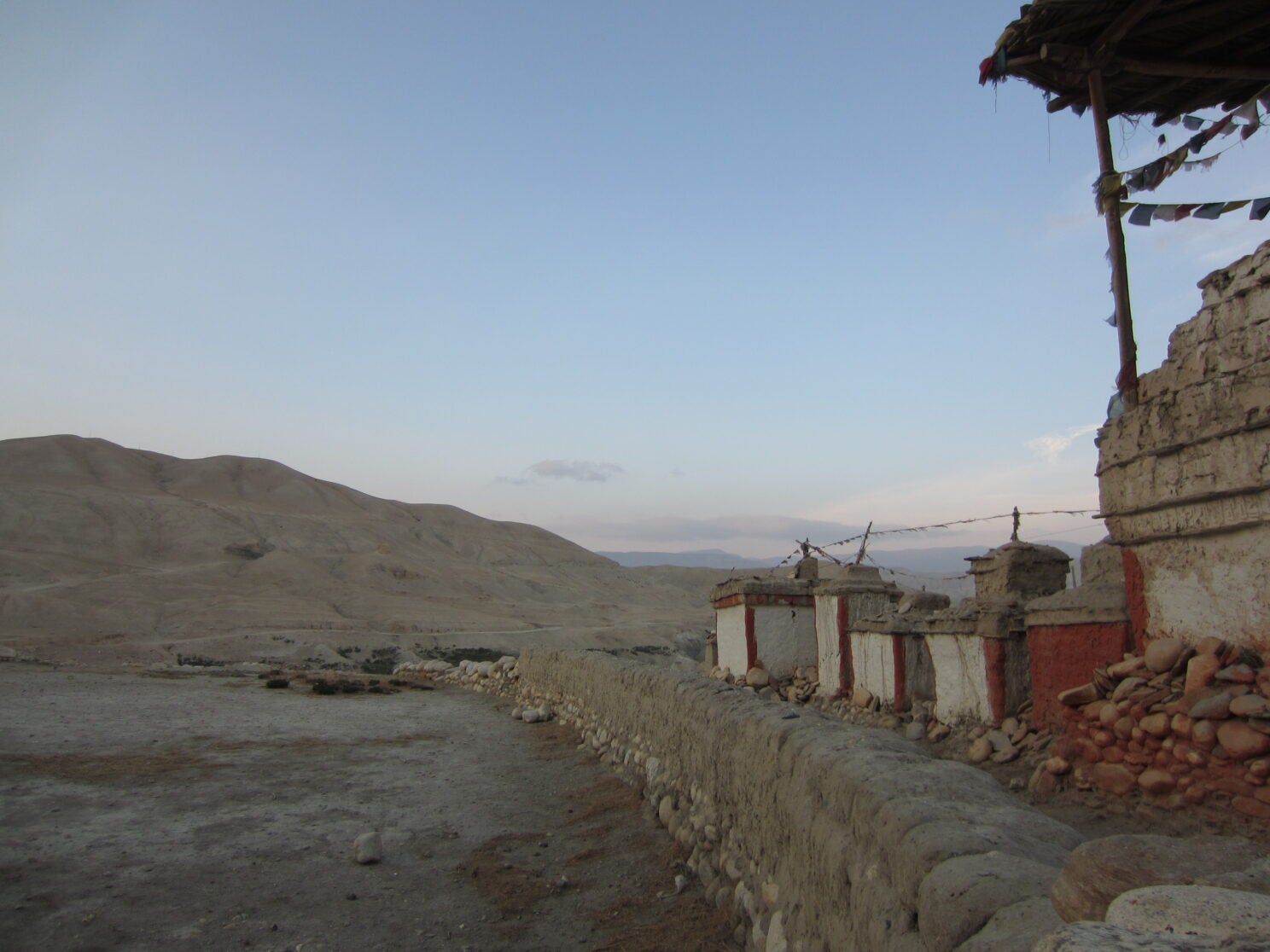

Upper Mustang region of Nepal.
Photo by Christina Warinner
New study of ancient genomes tracks disease over 5,500 years, factors in spread, including trade, warfare, colonialism, and slavery
Today, Malaria represents a major public health problem across much of the globe, killing more than 600,000 people annually and infecting another 250 million. It is a disease that has been around for millions of years and is undeniably entwined with human history.
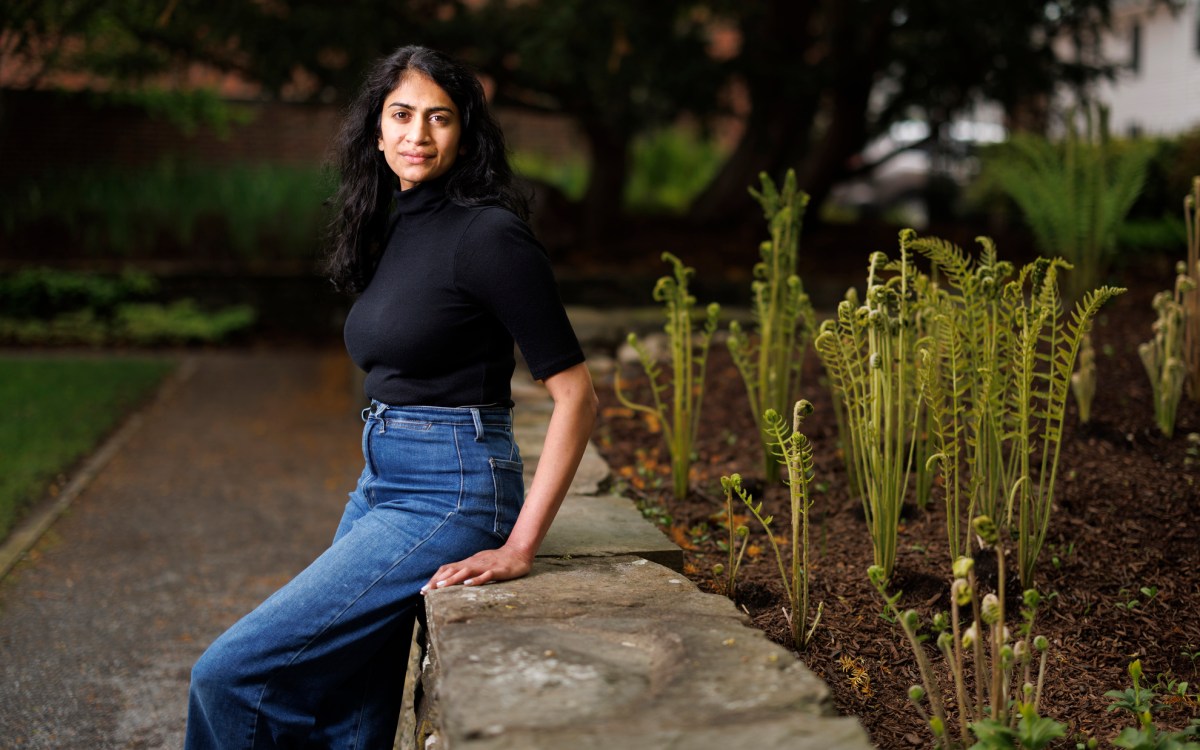
“Malaria has actually shaped the human genome,” said Megan Michel, a Ph.D. candidate in human evolutionary biology at the Harvard Kenneth C. Griffin Graduate School of Arts and Sciences. She pointed out that certain inherited blood disorders, like sickle cell disease, rose in prevalence because they provide a measure of resistance to the mosquito-borne infection.
Now a new study led by Michel reconstructs the ancient genomes of the two deadliest malaria parasites, Plasmodium falciparum and Plasmodium vivax, with an eye to understanding the pathogen’s past. The research, published this week in Nature, tracks the disease over 5,500 years, with trade, warfare, colonialism, and slavery identified as major factors in its global spread.
The findings represent a feat of collaboration and data-sharing, with 94 co-authors representing 80 institutions and 21 countries. The DNA itself was plucked from genetic sequences collected from more than 10,000 ancient humans, with Michel identifying 36 malaria-infected individuals from 26 archaeological sites on five continents.
“For a graduate student to be coordinating all of this is really, really impressive,” said co-author Christina Warinner, the John L. Loeb Associate Professor of the Social Sciences and one of Michel’s three advisers. “By reconstructing these ancient Plasmodium genomes and comparing the genetic relationships between ancient and modern parasites, we’re finally able to place malaria in its evolutionary and human history context.”
“By reconstructing these ancient Plasmodium genomes and comparing the genetic relationships between ancient and modern parasites, we’re finally able to place malaria in its evolutionary and human history context.”
Christina Warinner, the John L. Loeb Associate Professor of the Social Sciences
Malaria is marked by cyclical fevers that repeat every 48 or 72 hours. Until recently, written records were the only way researchers could track the disease’s progression across time and space. “There are descriptions in Greek and Roman texts that point to the presence of malaria,” Michel said. “But we were able to go back even further than that to show that malaria has been present in Europe for a very, very long time.”
The disease was also common in the U.S. until the arrival of modern drainage and insecticides in the 20th century. Warinner, a biomolecular archaeologist, pointed to the high number of U.S. presidents to suffer from malaria, including George Washington, Abraham Lincoln, and Ulysses S. Grant. “Teddy Roosevelt and JFK became infected while traveling,” she said, “but earlier presidents contracted it in their hometowns or in the Washington, D.C., area” — which was notoriously swampy.
The new paper features three compelling case studies, each illustrating the role of mobility in circulating malaria. The first concerns a Belgian cemetery, excavated between 2009 and 2011 and adjacent to the first permanent military hospital in early modern Europe. Historical records document that the Habsburg Army of Flanders recruited its soldiers from the Mediterranean region for its 80 Years’ War against Spain (1568-1648).
Malaria leaves no visible trace in human skeletal remains, but recent technological advances have enabled scientists to extract DNA from scraps of the pathogen found in teeth. Researchers were able to sequence malaria DNA from 10 individuals buried at the cemetery while also analyzing the genomes of soldiers who had been infected.
“We found that individuals buried at the cemetery have diverse ancestry profiles,” explained Michel, whose Ph.D. research is supported by the Max Planck - Harvard Research Center for the Archaeoscience of the Ancient Mediterranean. “They’re not just from Belgium. They seem to also be coming from northern Spain and from Italy.”
The two most prevalent malaria parasites in humans are P. vivax and P. falciparum, with the latter limited to warm climates and causing a more severe form of the disease. Analyses of pathogen DNA turned up a couple of P. vivax cases in the Belgian site’s local population, buried at the cemetery prior to the hospital’s construction in the mid-16th century.
Six cases, including several of the more virulent P. falciparum, were found in non-local individuals, all interred following the military hospital’s construction. Malaria cannot be transmitted through human contact, but mosquitos may have picked up these infections — and kept up the spread from there. “It’s even possible they ignited a local outbreak,” Michel said.

A bite from an infected mosquito transmits malaria.
Liz Zonarich/Harvard Staff; source: Mayo Clinic
The parasites travel to your liver where they lie dormant, usually about 10 days to four weeks.

Parasites leave the liver and infect red blood cells. Malaria signs and symptoms typically develop.

Malaria is transmitted to an uninfected mosquito when it bites someone with the disease.
Another case study came from Peru, where a single P. vivax case was found in a person who lived at high altitude (more than 9,300 feet) in the Central Andes. “This individual was associated with the Chachapoya culture,” Michel said, “and the site we were working with spanned the period of European contact.”
For years, scientists have debated how the disease arrived in the Americas, where Indigenous populations lack genetic resistance to malaria. Reconstructing the genome of the Peruvian parasite revealed striking similarities to P. vivax strains found throughout South America today. It also resembled strains circulating in Europe during the 15th and 16th centuries.
“We think this is evidence that the species was transmitted by European colonizers to the Americas,” Michel said.
No ancient P. falciparum was found in the Americas, Michel noted, and P. falciparum strains circulating there today bear little resemblance to the ancient European P. falciparum parasites recovered by Michel and her co-authors. “Instead, American strains today look very similar to strains in Sub-Saharan Africa,” Michel said. “It seems likely that P. vivax was transmitted from Europe, whereas P. falciparum was probably transmitted from Sub-Saharan Africa as a result of the trans-Atlantic slave trade.”
Michel got her biggest surprise from the paper’s third case study. The Himalayan site of Chokhopani, situated more than 9,100 feet above sea level in Nepal’s Mustang region, yielded the earliest known case of P. falciparum.
“It’s the last place on Earth I would expect to find a malaria infection,” Michel shared. “It’s rocky and dry and too cold for malaria-transmitting mosquitoes to survive.”
The infected individual lived 2,800 years ago. “We know from the archaeological record that there was extensive long-distance trade in the region,” explained Michel, who partnered with co-author Mark Aldenderfer — an archaeologist working in the Mustang region for many years — to analyze the findings. “We think this was probably an individual who moved from low to high altitude, possibly for trade. They must have acquired this infection at a lower altitude where the parasite can be transmitted.”
“The site of Chokhopani is near the Kora La pass, the lowest crossing point through the Himalayas and a key trade route connecting South Asia with the Tibetan Plateau,” added Warinner, who traveled with Michel to the region last spring to share results and solicit feedback from descendent communities. “Fortunately, Nepal has been really successful in eradicating malaria in the last few years. But even as recently as 10 years ago, malaria was endemic in Nepal’s lower elevation regions.”
Making these revelations possible are the emerging tools of metagenomics, which rely on recovering and sharing as much genetic data as possible with different specialists. “When we analyze an ancient sample, we, by its nature, destroy it in order to retrieve the DNA,” Warinner explained. “We want to get as much information as possible. We really do recover total DNA.”
“Metagenomics and data-sharing allow us to find things we’re not really looking for,” Michel added. “It lets us find disease in unexpected places. I never would have screened samples from Chokhopani for malaria if they hadn’t already been sequenced by Dr. Warinner for another ancient DNA study.”
The research described in this report received funding from the National Science Foundation




