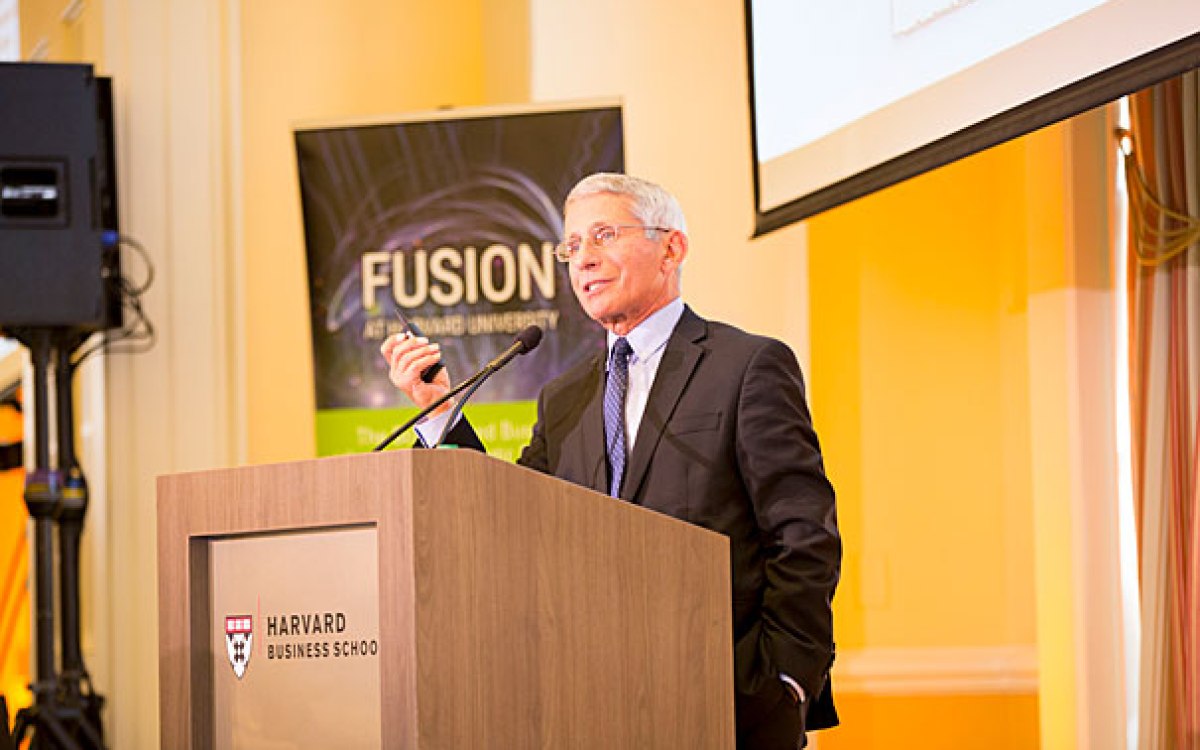
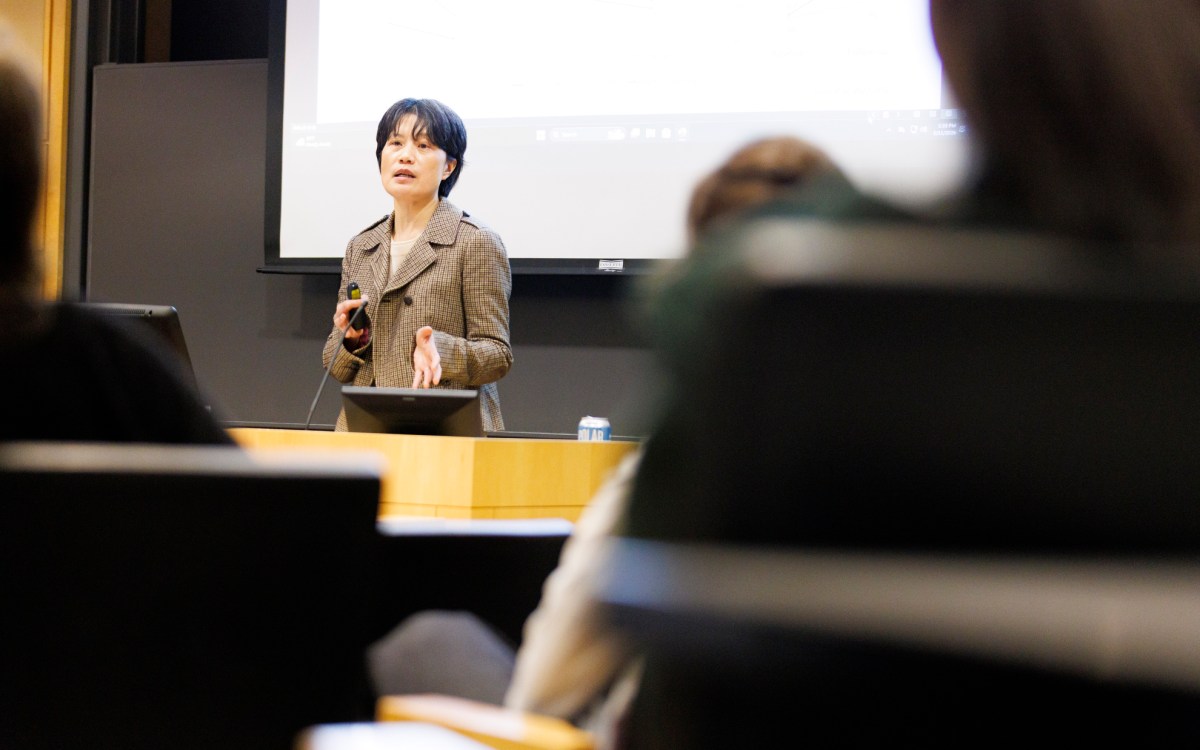
Jim Collins opens “Data-Powered Strategies to Counteract Antibiotic Resistance,” a Medical School conference on how big data can be utilized to fight antibiotic resistance.
Rose Lincoln/Harvard Staff Photographer
Big data targets drug resistance
Harvard symposium examines bioinformatics’ great potential in ensuring antibiotics’ effectiveness
Experts who are adept at handling massive scientific data sets are examining ways that the revolution sweeping their field can address one of medicine’s biggest challenges: the spread of drug-resistant microbes.
Maha Farhat, assistant professor of biomedical informatics and one of the organizers of a symposium on the topic, said antibiotic resistance has reached “epidemic proportions” but, with more data available than ever, there’s hope that medical science will discover ways to combat it. What’s needed, however, are better tools to analyze that data, she said.
“It really threatens one of the most core therapies that define modern medicine,” Farhat said. “We do think the problem of antibiotic resistance is at such scale that we should come at it from many different angles.”
The symposium, “Data-Powered Strategies to Counteract Antibiotic Resistance,” held Wednesday at Harvard Medical School’s New Research Building on the Longwood campus, featured speakers who focused on three major areas: detecting and diagnosing drug-resistant microbes, applying big data to the problem, and developing new drugs to fight resistance.
Speakers came from both academia and industry and represented Harvard, Massachusetts Institute of Technology, Imperial College London, McMaster University, Boston Children’s Hospital, Entasis Therapeutics, and Accelerate Diagnostics.
Harvard Medical School Dean George Daley introduced the session, calling the years after penicillin first became widely used in the 1940s a time of relative innocence. Penicillin was so effective in fighting infections that previously would have been fatal that it transformed not just the hospital, but the battlefield.
The rise of drug resistance in the decades since, however, has resulted in some 2 million drug-resistant infections that kill 23,000 people in the U.S. each year. The resistance also costs the medical industry billions a year in higher medical costs.
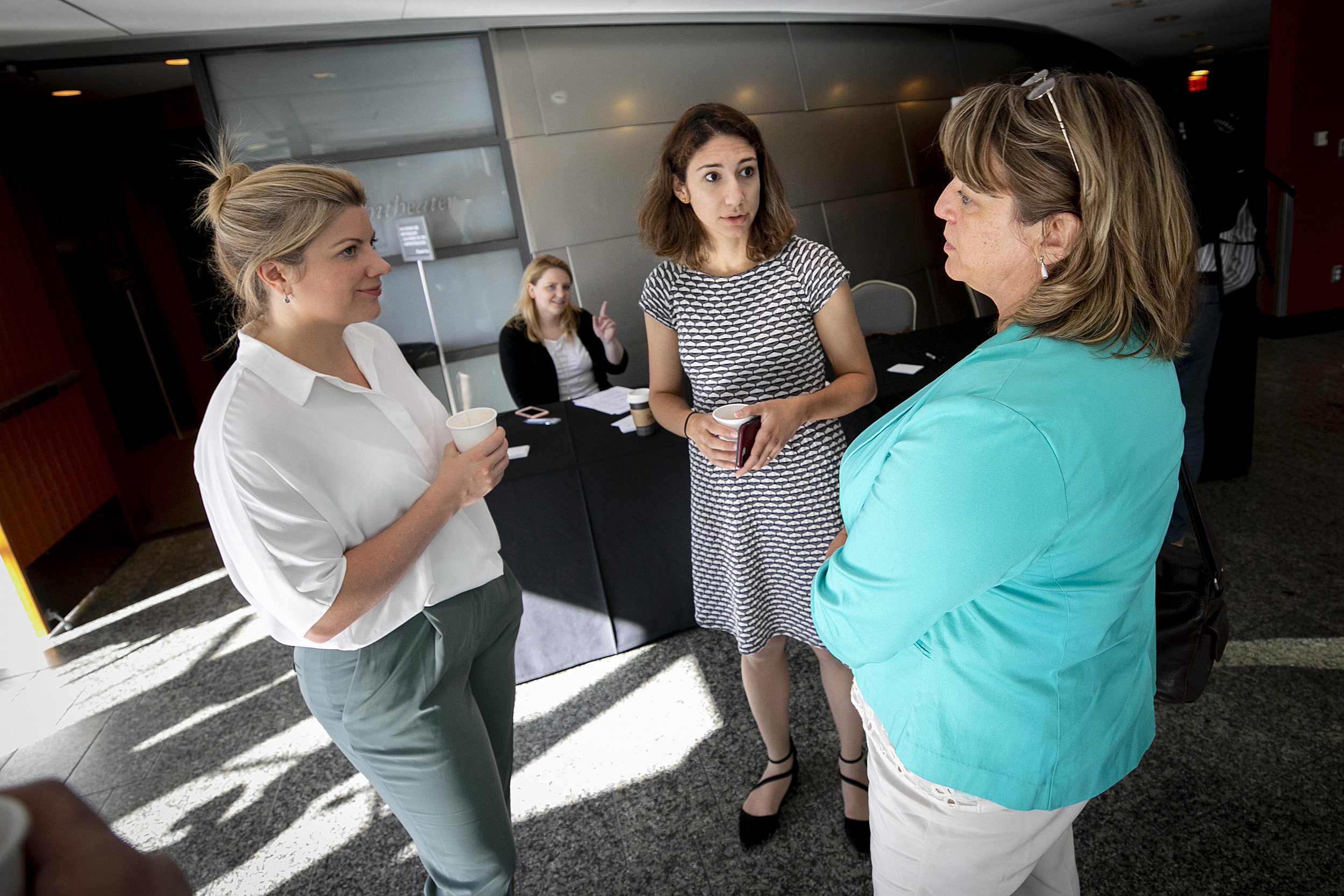

Conference organizer Maha Farhat (center) talks with speakers Ceire Costelloe (from left) and Alita Miller.
Rose Lincoln/Harvard Staff Photographer
Daley, who, before becoming dean directed the pediatric bone marrow program at Boston Children’s Hospital and the Dana-Farber Cancer Institute, said physicians there — concerned about infection because bone marrow transplants weaken the immune system — could see that the tools they had at their disposal were weakening.
“The fact is that the diminishing potency of our prophylactic armamentarium really was quite frightening. There were lots of diseases and infections that, once they set in, despite our efforts at prophylaxis, it became very, very challenging,” Daley said. “In virtually every medical context, the physicians’ arsenal of antimicrobial therapies has become a rapidly dwindling set. New therapies are just not being developed fast enough.”
The problem is alarming enough, Daley said, that in 2016 the World Health Organization declared it one of the greatest threats to global health and made it a top priority.
Though improper use of drugs — for the wrong microbes, or in regimens that don’t completely eradicate the infection, leaving the hardiest surviving microbes alive to multiply — can increase resistance, it’s also a natural process, part of an ongoing arms race between microbe and host, Daley said. Research has found mechanisms of drug resistance in 30,000-year-old microbes frozen in permafrost, indicating that it is a natural phenomenon and not a result of poor prescribing practices and patient adherence.
“What it says is that the constant battle between pathogen and host … has been an ancient battle. It’s now playing out in the human body,” Daley said.
While Daley said that new antibiotic drugs are needed, progress in fighting resistance will come from other sources as well, including leveraging insights about bacteria gleaned by analyzing the large data sets newly available. One potentially fruitful area, Daley said, is the vast amount of genomic data now available that can provide a better understanding of a microbe’s weaknesses.
Thousands of tuberculosis genomes have been analyzed, for example, as researchers look to better understand patterns of drug resistance in a bacterium that ranks high on a global list of the most worrisome drug-resistant infections.
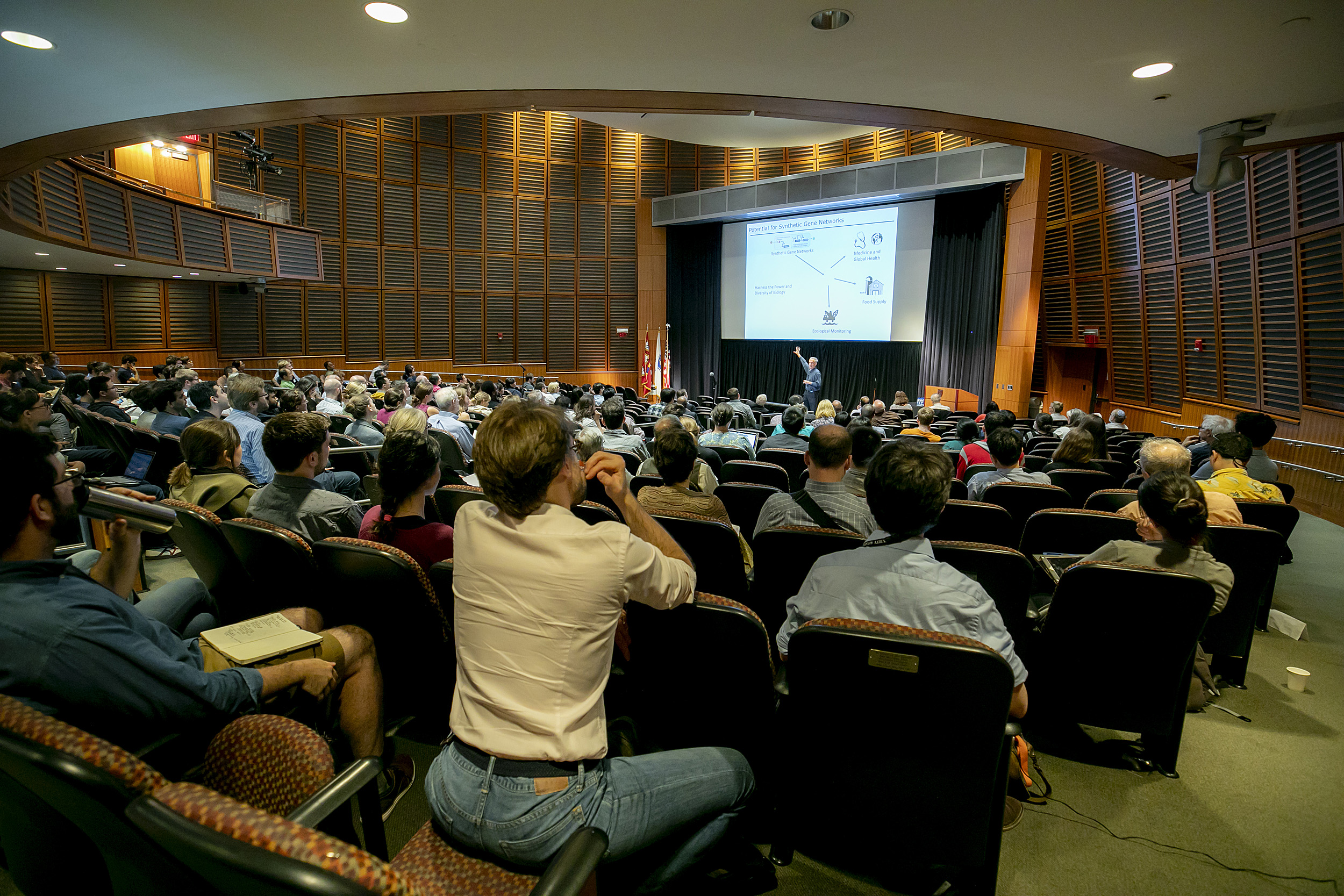

A near capacity crowd filled the lecture hall at HMS’ New Research Building.
Rose Lincoln/Harvard Staff Photographer
One important area where progress is being made is in rapid diagnostics. A severely ill patient suffering from an infection is usually treated right away, without waiting for lab results to provide information about a particular strain and its susceptibility to antibiotics, which can take days. James Collins of MIT, the Broad Institute of MIT and Harvard, and Harvard’s Wyss Institute for Biologically Inspired Engineering, and Romney Humphries of Accelerate Diagnostics presented two early talks about rapid diagnostics.
Humphries described her company’s phenotype-based test, which cuts diagnosis time from several days to about seven hours. That’s still too late to inform the first application of antibiotics, Humphries said, but could allow physicians to shift medication strategies for second and subsequent doses should a strain be determined to be resistant to particular drugs. What’s needed, she said, are diagnostics available within 30 minutes, so physicians can avoid prescribing inappropriate or ineffective medicines from the start, which not only do the patients no good, but risk enhancing resistance.
Collins, the Termeer Professor of Medical Engineering and Science at MIT and a founding core faculty member at the Wyss, described the development of engineered bacteria that function as living diagnostics, and of inexpensive, paper-based diagnostics that can be used in remote locations.
He also described a new approach to fighting infection by harnessing the microbiome. Researchers in his lab seeded the intestines of mice with a microbe found in dairy products, one that they’d engineered to detect cholera and that in its natural form makes the gut more acidic. That makes the gut a more hostile environment for the cholera bacterium, staving off its potentially deadly symptoms.
“This opens up a broad range of possibilities,” Collins said. “This is something you can have in a simple pill or literally spike in yogurt or fermented dairy products” when a pandemic threatens.