Whole brain imaging
Lichtman-led zebrafish findings open path to deeper insight on behavior
Scientists appear closer than ever to unlocking the black box that is the brain, and they’re doing it with the help of a fish less than half an inch long.
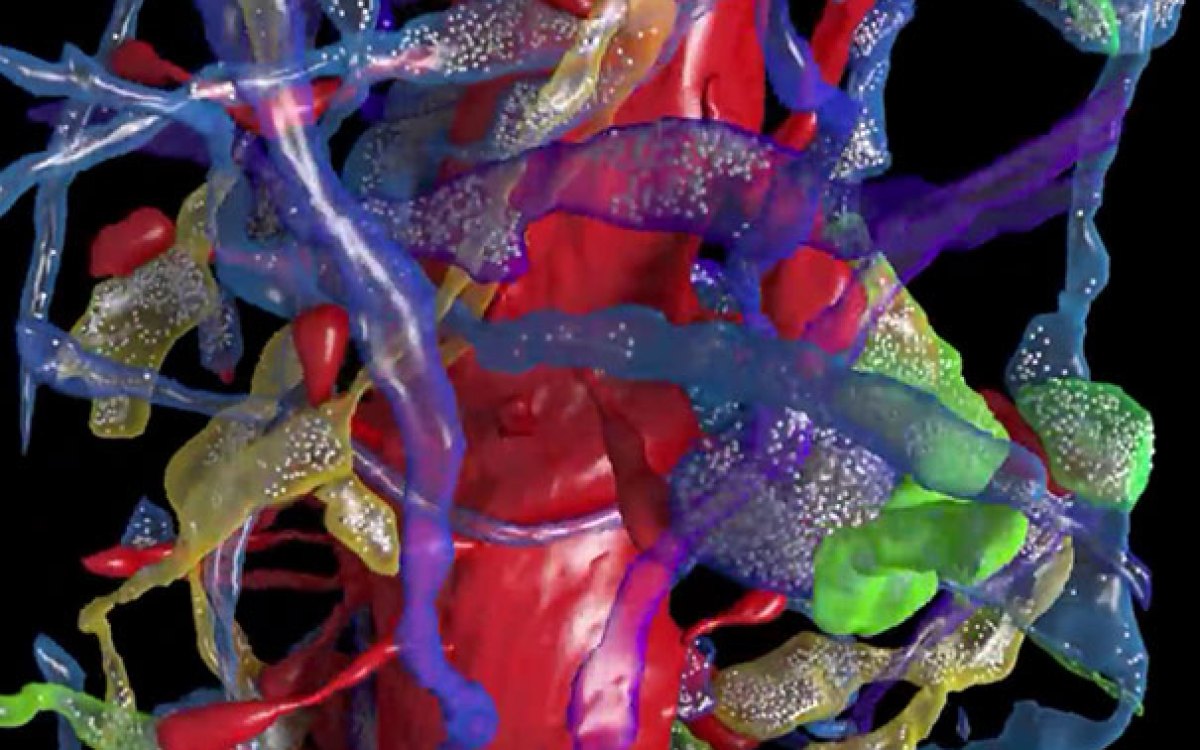
Led by Jeff Lichtman, the Jeremy R. Knowles Professor of Molecular and Cellular Biology and Santiago Ramon y Cajal Professor of Arts and Sciences, Florian Engert, a professor of molecular and cellular biology, and David Hildebrand, a postdoctoral fellow in Engert’s lab, a team of researchers has imaged every neuron in the brain of a larval zebrafish at nanoscale resolution.
The breakthrough, described in a May paper in the journal Nature, will allow detailed investigation of how stimuli are processed by the brain and give rise to specific behaviors, the researchers said.
Fundamentally speaking, Hildebrand said, the study serves as a proof of concept for a new approach to nanoscale brain imaging — by making some sacrifices in resolution, the team was able to image the entire brain, leaving open the possibility of scanning specific areas of interest with greater resolution.
“Most studies have focused on imaging a small section of brain with very high resolution,” he said. “Instead of simply imaging a small section, we’ve actually pushed the methodology … so we can capture the entire brain of the zebrafish, and then as our questions require it, image regions at higher and higher resolution.”
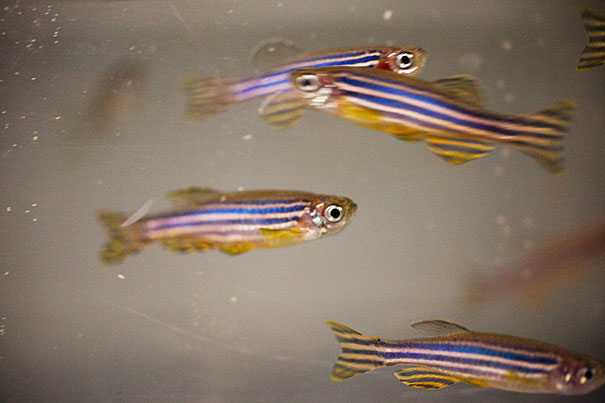

But even at lower resolution, Hildebrand said, interesting details jumped out.
As part of the study, he said, the team uncovered an unusual level of symmetry among myelinated neurons — nerve fibers encased in an insulating layer, enabling signals to travel faster.
“When we zoomed in on one particular subset, we found their position and shape were strikingly symmetrical across the midline of the fish,” Hildebrand said. “This [might] give us some insight into what type of developmental mechanisms control how their shapes are established. It may even allow us to start thinking about how these mechanisms may influence their connectivity to downstream targets.”
“When you do something like this in a mammal … we found a long time ago, there’s no symmetry at all,” Lichtman added. “And not only that, the wiring diagram in that case is largely evolved during development through mechanisms that shape the nervous system to match the world it finds itself in. So it’s possible that these fish neurons that show this kind of stereotypy of left and right are wired based on nature rather than nurture.”

In addition to probing the developmental mechanisms behind how the brain is wired, the study points to a new method for understanding the neural circuits that drive certain behaviors.
To do that, Engert said, researchers start with a precise description of a particular behavior. Using that description and known anatomy, they can develop hypotheses of how the brain is processing sensory information to generate behavior. Using data collected from ultrastructural brain imaging, they can then test whether the assumptions about the anatomy underlying those models are accurate.
“But even more importantly, they can be used to reject hypotheses,” Engert said. “If what you find invalidates your model … then you’ve made progress in a very dramatic and painful way, but science advances when it hurts.”
“One of the limits of imaging small chunks of brain is that it’s harder to infer what the inputs and outputs of the system are,” Hildebrand added. “Since we have essentially the entire brain of this organism, we can see the sensory inputs directly … and we can go all the way through that circuit and get back to some muscle group. It’s an unprecedented level of completeness.”
More like this
The researchers share another goal, Engert said — to make these data sets available to the scientific community as a sort of “whole-brain atlas,” which can be used to address any number of questions.
“A good analogy is the sequencing of the genome,” he said. “Initially, that was a huge effort, and the next mountain to climb was to apply that technology to scientific questions. That’s one of the main goals here. We hope this can become a more routine tool for research … as a technology to compare brain structure and function.”
The research was supported with funding from National Institute of Health, the Defense Advanced Research Projects Agency, the National Research Foundation of Korea, and the National Science Foundation.