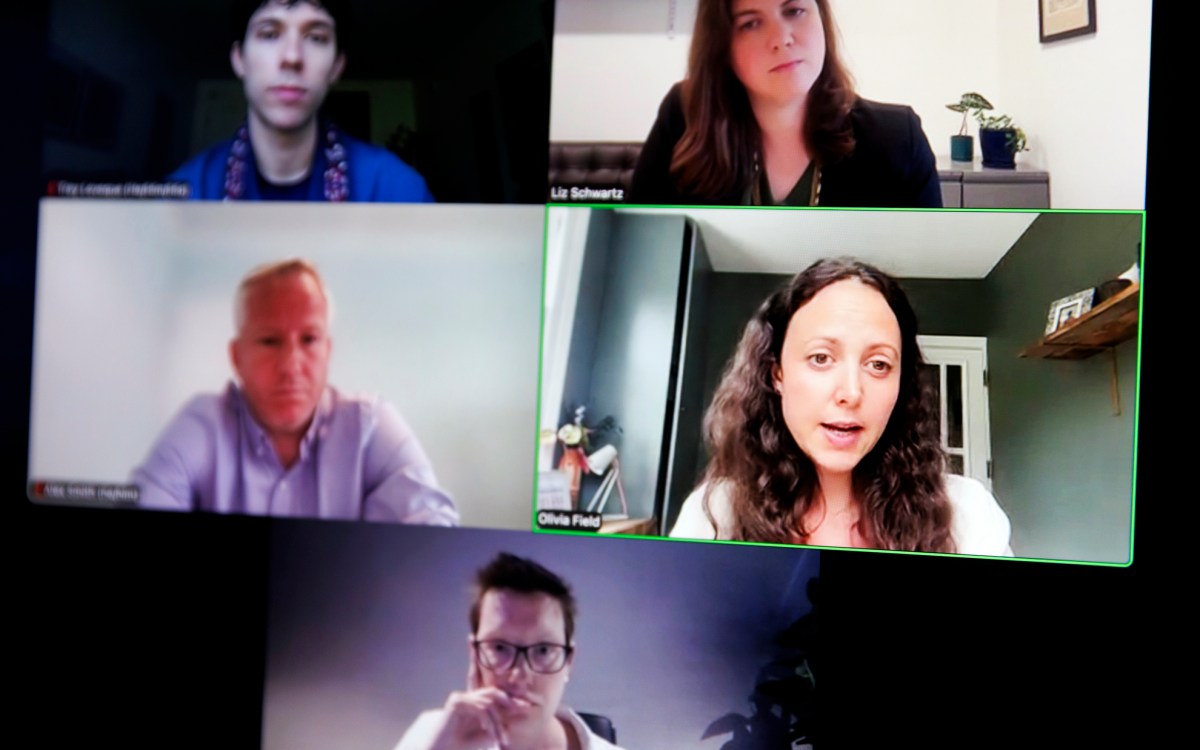
Using genetic tools, researchers at Harvard and collaborating institutions have completed the most comprehensive evolutionary tree ever produced for mollusks. Pictured is a bivalve collected in Moreton Bay on the eastern coast of Australia. In a scientific first, the results suggest that bivalves, a class of animals that includes clams and oysters, are most closely related to gastropods such as snails and slugs.
Image courtesy of Gonzalo Giribet
Mapping mollusks
Researchers use genetic tools to complete family tree
What do a typical garden snail and an octopus have in common, besides the occasional appearance on the plates of adventurous diners? More than you may realize. Both are mollusks, a group of animals that includes everything from slugs to squids, and one that has for years confounded efforts to describe how its varied members evolved.
But now, with the help of the latest genetic tools, researchers at Harvard and collaborating institutions have completed the most comprehensive evolutionary tree ever produced for mollusks. Described in the Nov. 2 issue of Nature, the work also serves as a proof-of-concept, demonstrating the power of genomic techniques to answer difficult evolutionary questions.
“I think this is a big step toward understanding the tree of life,” said Gonzalo Giribet, professor of organismic and evolutionary biology, and one of three lead authors of the paper. “One reason we chose mollusks was because it has been one of the hardest phyla to reconstruct phylogenetically. This shows very clearly that we can now use this technique to resolve other evolutionary problems.”
Using more than a dozen mollusk species collected from around the world, Giribet and other researchers went to work, sequencing nearly every gene in the animal before selecting 1,800 specific genes to compare across species. By comparing those sequences with each other, and with genetic information from other species, they were able to reconstruct the ways in which all mollusk species are related to one another.
In a scientific first, the results suggest that bivalves, a class of animals that includes clams and oysters, are most closely related to gastropods such as snails and slugs. Other findings, Giribet said, resolve the divergence between mollusks that developed shells and those that did not.
Perhaps the paper’s most interesting finding, however, is that monoplacophorans, a type of mollusk thought to be extinct until the early 1950s, are most closely related to cephalopods, a group that includes octopus and squid. Similar theories had been advanced by paleontologists based on the fossil record, Giribet said, but, until now, without the support of biological analysis.
Understanding the phylogeny of an animal, Giribet said, is a critical first step in understanding how developmental mechanisms and anatomical changes evolve over millions of years.
“The first step is in knowing the relationship between animals,” Giribet said. “We need to have that phylogenetic framework, because otherwise we’re making the wrong comparisons.
“At one point, we thought we would never be able to resolve something like the Cambrian explosion, because there was simply too much diversity appearing in too short a time, but one of the reasons for that was that we had very poorly resolved phylogenies,” Giribet continued. “What we’re showing is that by examining not one or two or 10 or 50 genes, but by looking at hundreds or even thousands of genes we are able to reconstruct any evolutionary event that happened during early animal evolution.”
The other lead authors who collaborated on the research were Casey Dunn, an evolutionary biologist at Brown University, and Nerida Wilson from the Australian Museum. Sónia Andrade, a former Harvard postdoctoral student working in Giribet’s lab, also contributed to the paper.
The research was funded by the National Science Foundation, Scripps Institution of Oceanography, the University of California Ship Funds, the Harvard Museum of Comparative Zoology, and the Carlsberg Foundation.